|
Modeling and anlysis of biological networks.
The complexity of a living cell is controlled by several genes and their products, or sets of co-regulated genes sharing a common function. With the availability of complete genome sequences and high-throughput post-genomics experimental data, it is possible to study networks of macromolecular interactions such as gene regulatory networks (GRNs), metabolic networks, and protein-protein interaction (PPI) networks. Computational modeling and anlysis of biological systems has become important in Systems Biology research in order to understand the complex biological interactions and behaviour. After several years of research, most of the interactions and parameters of the cellular networks are still not known or poorly understood. Hence, accurately modeling such systems still remains a challenge for the researchers. |
|
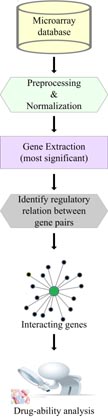
Analysis of microarray and next-generation sequencing data
From the past several decades, microarray technology has been extensively applied as an indispensable tools to simultaneously monitor gene expression on a genome-wide scale. This technology has been widely used for the study of cancer genomes and transcriptome but some its promises have not materialized due to limitations of these data. With the development of Next-Generation Sequencing (NGS) technology, it has surpassed and replaced microarray technology allowing precise analysis of RNA transcripts for gene expression study. The NGS has also replaced the limitation of microarrays to detect poorly expressed genes. It is also used as a tool for the identification and analysis of DNA regions that interact with regulatory proteins in functional regulation of gene expression. NGS offer novel, rapid ways for genome-wide characterisation and profiling of mRNAs, small RNAs, transcription factor regions, chromatin structure and DNA methylation patterns.
In this direction, we are working on the development of intelligent algorithm for better analysis of microarray as well as NGS data.
 +91-11-26980014 Extn. 30
+91-11-26980014 Extn. 30 
 /
/





 Home
Home